Menu
D3AI-Spike
Introduction
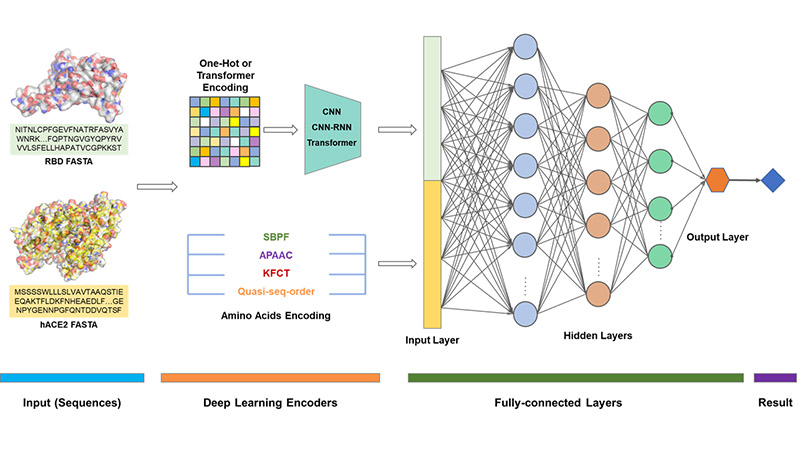
Under the wreak havoc of SARS-CoV-2 since the end of 2019, the epidemic of COVID-19 has had a tremendously serious effect on mankind. Although researchers have been developing various drugs and vaccines fighting against COVID-19 round the clock, various mutants have different caused immune escapes, which brings great more challenges to us. But the test cycle for evaluating each new variant is relatively long, which generally takes several weeks. The infectivity and severity of the SARS-CoV-2 variant largely depend on the affinity of its S protein receptor-binding domain (RBD) and hACE2. Therefore, based on the affinity data between RBD and hACE2 and broad-spectrum proteins, we developed an open network platform based on several AI prediction models to facilitate researchers all over the world predicting the binding affinity changes of new mutant strains in a very short time, which provides a first-time reference. Different AI methods are used to predict their affinity and the relatively accurate prediction models are selected then comprehensively analyzed to get the final result.
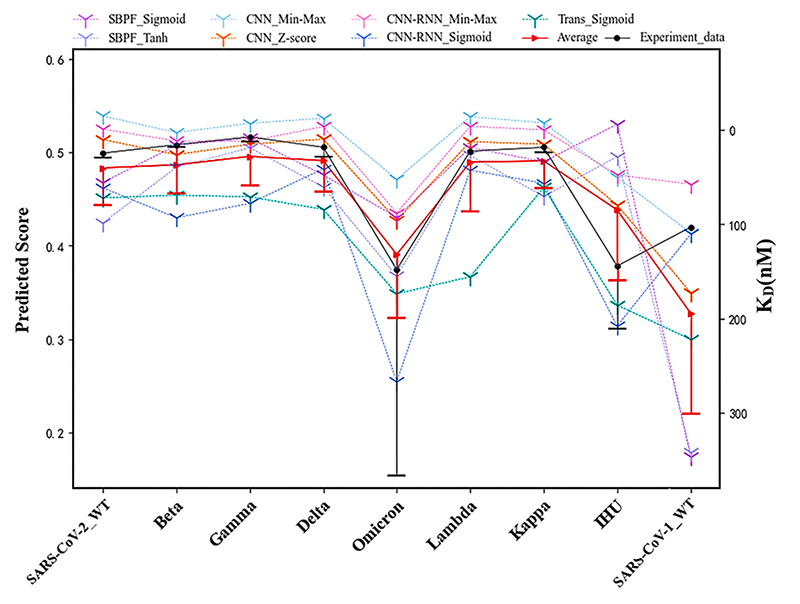
Several variants are predicted to compare the binding affinity with SARS-CoV-2-WT. The black and red error bars represent the confidence intervals of experimental and predicted values.
This site has been visited 4595346 times since Feb 2020.