Menu
D3DistalMutation ( >10Å )
DistalMutation ( >10Å )
Mutation ( <10Å )
Welcome to D3DistalMutation database
In general, D3DistalMutation describes the effect of distal mutation (mutations more than 10 Å away from the active site) on enzyme activity and classified enzymes into four classes, viz., no activity change, decrease activity, loss of activity and increase activity. The D3DistalMutation provides information including mutation residues, active sites, the closest distances, activity change description and references. We hope that the D3DistalMutation can facilitate allosteric drug design and industrial catalysis research.
Downlaod the code and data to reproduce D3DistalMutation. It has been downloaded by 817 users .
Searchable:
Users can make global search and fuzzy search of D3DistalMutation.
Downloadable:
Users can click entries under the title of “Enzyme name” to download three-dimensional pse files of D3DistalMutation. The three-dimensional pse files highlighting mutant residue, active residues, and predicted pockets for further research could be visualized by PyMOL. In the pse files, pockets sequence numbers in ascending order correspond to pockets volume in descending order, for example, the poceket1 house the largest pocket volume.
Downlaod the data of D3DistalMutation. It has been downloaded by 584 users .
Linkable:
Users can click entries under the title of “Uniprot ID”, “PDB ID” and “References” to jump to UniProt, PDB and PubMed, respectively.
Statistics:
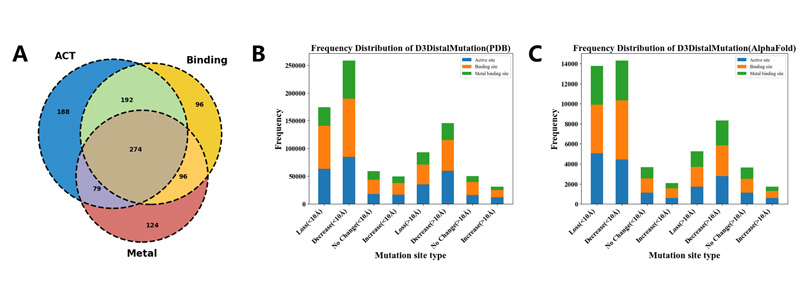
Based on different protein activity changes in different functional sites (Active Site, Binding Site and Metal Binding Site).The D3DistalMutation 2.0 database contains 7201 proteins from 1049 species, which were classified into four categories (no activity change, decreased activity, loss of activity, or increased activity). The updated database has been expanded by 244% compared to the previous data of 2130 proteins.
 |
